This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison
Domains in CDH23 Homo sapien protein
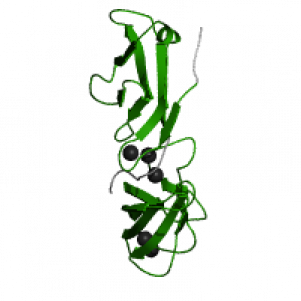
Background info on the Cadherin domain
- GO process:homophilic cell adhesion (GO:0007156)
- GO component:membrane (GO:0016020)
- GO function:calcium ion binding (GO:0005509)
These will be discussed in detail in the protein ontology page. (EMBL-EBI GO Finder)
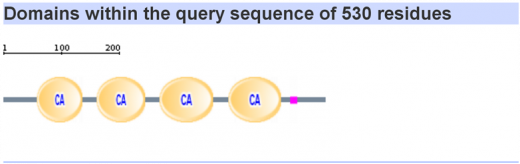
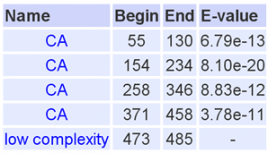
Domains in the Homo sapien cadherin-like 23 protein
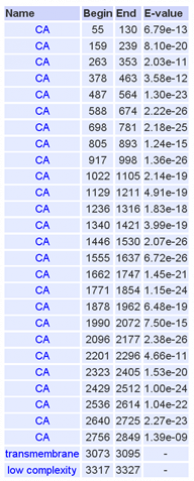
A total of 26 cadherin repeat domains were found within the human cadherin-like 23 protein that we have been using to compare against other model organisms. Using hydrophobic analysis a transmembrane segment was also found at the end of the sequence. This may indicate the glyco- attachment site of this glycoprotein, suggesting that all cadherin repeat domains of the protein are localized to the extracellular space in vivo.
All detected cadherin repeats met the significance threshold (stated above) for attaining significance as a domain.
Below we have analyzed 4 isoforms of this particular protein, to determine which, if any, domains are missing.
Protein interactions within cdh23
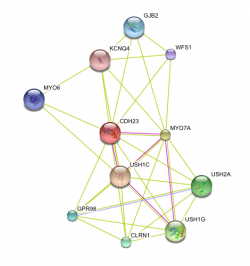
A network interaction between cadherin 23 and other related proteins was made using STRING. USH1C, or harmonin, has been identified as a potential partner in function with CDH23. USH1C is known as the "Usher Syndrome type 1-C" protein, because its loss of function is known to cuase this subtype. USH1C is suggested to be involved in protein-protein interaction.
I would suggest that studies investigating the possible function of USH1C in binding mutiple cells' CDH23 together be undergone. See protein network interactions page for more info.
I would suggest that studies investigating the possible function of USH1C in binding mutiple cells' CDH23 together be undergone. See protein network interactions page for more info.
Isoforms of CDH23 Protein and Domain Variances
We used the SMART domain identifier program to further analyze discrepancies in domains between wild type cdh23 and isoforms. We were able to find discrepancies in cadherin domain numbers among isoforms:
-Isoform 3: 12 cadherin domains
-Isoform 4: 9 cadherin domains
-Isoform 5: 3 cadherin domains
-Isoform 6: 5 domains
Further research could identify specific cadherin domains missing from the original protein, and a comparison of pathogenic isoforms could shed light on whether a domain is necessary for normal function, or if a certain range of domains seem to be correlated with a pathogenic defect.
-Isoform 3: 12 cadherin domains
-Isoform 4: 9 cadherin domains
-Isoform 5: 3 cadherin domains
-Isoform 6: 5 domains
Further research could identify specific cadherin domains missing from the original protein, and a comparison of pathogenic isoforms could shed light on whether a domain is necessary for normal function, or if a certain range of domains seem to be correlated with a pathogenic defect.
References:
SMART: http://smart.embl-heidelberg.de/
Expasy: http://ca.expasy.org/prosite/
EMBL-EBI GO finder: http://www.ebi.ac.uk/QuickGO/GProtein?ac=A5D6V9
String: http://string-db.org/newstring_cgi/show_input_page.pl?UserId=122XyLCvXqZ_&sessionId=iqPHGxezn071
SMART: http://smart.embl-heidelberg.de/
Expasy: http://ca.expasy.org/prosite/
EMBL-EBI GO finder: http://www.ebi.ac.uk/QuickGO/GProtein?ac=A5D6V9
String: http://string-db.org/newstring_cgi/show_input_page.pl?UserId=122XyLCvXqZ_&sessionId=iqPHGxezn071
Ben Hofeld, [email protected], last updated: 5.15.2010, Link to course page:www.gen677.weebly.com